Imaging Brain State Dynamics
The main goal is to infer the spatiotemporal dynamics underlying cognitive processes. These dynamics are studied using different experimental techniques (TMS, MEG, EEG, fMRI). Of particular interest is to study the dynamical changes of large scale functional networks while executing a cognitive task or during resting conditions. Most of current methods to study these networks however do not take account of the fact that brain activity is highly dynamic. This limits how we investigate the healthy and the dysfunctional brain.
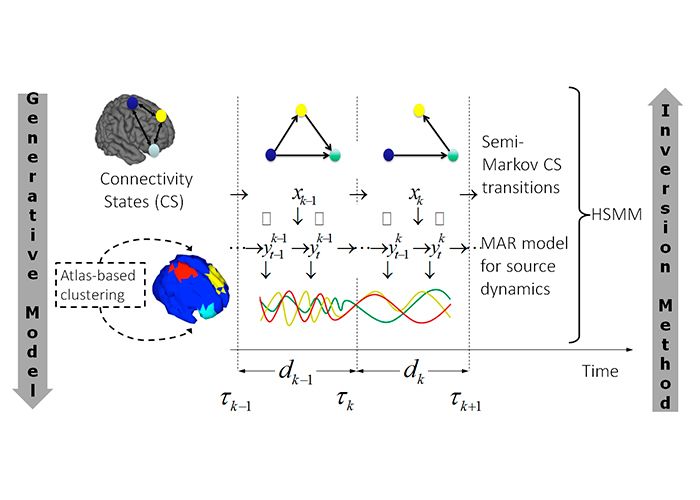
More specifically brain dysfunction may not be apparent until structural changes are visible or quantifiable (e.g. neuropsychological and clinical measures), which prevents effective and early intervention to target dysfunctions. Here we consider the brain as a stochastic dynamical system that can transition between Brain States (BS) or modes of operation and while in each mode, associated dynamics determine the evolution of the measured brain signals. Here, we consider BSs to represent functional brain networks that dynamically evolve over time. Then, we aim to develop novel methods to identify different types of BS involved in a given process and their dynamic evolution and to provide biophysically interpretable parameters based on the measured data.
These methods can then be used as tools for the early and non-invasive detection of abnormal operations of the brain, before plastic or structural changes become permanent. In addition, their parameters could be used as predictors in persons who will likely develop neurodegenerative diseases in their lifetimes.
Neurophysics
The Neurophysics theme aims at modelling the activity of the brain both in the normal and in the pathological case. For this purpose, we adopt a statistical physics approach to bridge different spatial and temporal scales of brain activity.
The ultimate goal is to derive models of large ensembles of neurons, which are measurable using non-invasive Neuroimaging techniques, while keeping neurobiological realism and interpretable parameters. To this end we use techniques from statistical physics to derive such large scale models based on realistic models of single neurons.
Of particular interest is the development of macroscopic models of brain plasticity to explain the relationship between plastic changes and neurotransmitter dynamics and how these changes can be detected using M/EEG, fMRI and MRS; and induced or modulated using neuro-stimulation techniques (TMS, TDCS, TCAS or sensory stimulation).
Computational models of behaviour
This line adopts a modern view of the brain as a Bayesian controller to explain behaviour.
This view is based on ideas from theoretical physics and the notion that the brain can be understood as an inference machine actively pursuing to minimise variational free energy.
To this end, the brain minimises free energy by modifying its configuration to change the way it samples the environment, or to change its expectations. These changes then correspond to behavioural processes such as action, perception and learning.
This formulation implies that the state and structure of the brain encode a probabilistic model of the environment that is continuously adapted through learning, and used to make predictions and evaluate prediction errors. Importantly the set of possible models is assumed to be fixed. We are interested in extending this framework to explicitly account for the possibility of learning a completely new (outside the set) representation of the environment. This entails allowing for considering an unbounded set of models under the constraint of limited brain resources. This will allow designing experiments to investigate a fundamental question with potential translational (diagnostic and therapeutic) implications:
How do brain mechanisms differ when we adapt a current model, switch to another already existing one, or learn a completely new model?